#include "MLIR/Analysis/ActivityAnnotations.h"

Public Member Functions | |
| void | print (raw_ostream &os) const override |
| ChangeResult | markAllOriginsUnknown () |
| const ValueOriginSet & | getOrigins (DistinctAttr id) const |
| ChangeResult | meet (const AbstractDenseLattice &other) override |
 Public Member Functions inherited from mlir::enzyme::MapOfSetsLattice< DistinctAttr, OriginAttr > Public Member Functions inherited from mlir::enzyme::MapOfSetsLattice< DistinctAttr, OriginAttr > | |
| Attribute | serialize (MLIRContext *ctx) const |
| ChangeResult | join (const AbstractDenseLattice &other) |
| ChangeResult | insert (const SetLattice< DistinctAttr > &keysToUpdate, const SetLattice< OriginAttr > &values) |
| Map all keys to all values. | |
| ChangeResult | markAllUnknown () |
| const SetLattice< OriginAttr > & | lookup (DistinctAttr key) const |
Additional Inherited Members | |
 Protected Member Functions inherited from mlir::enzyme::MapOfSetsLattice< DistinctAttr, OriginAttr > Protected Member Functions inherited from mlir::enzyme::MapOfSetsLattice< DistinctAttr, OriginAttr > | |
| ChangeResult | joinPotentiallyMissing (DistinctAttr key, const SetLattice< OriginAttr > &value) |
 Protected Attributes inherited from mlir::enzyme::MapOfSetsLattice< DistinctAttr, OriginAttr > Protected Attributes inherited from mlir::enzyme::MapOfSetsLattice< DistinctAttr, OriginAttr > | |
| DenseMap< DistinctAttr, SetLattice< OriginAttr > > | map |
| Maps a key to a set of values. | |
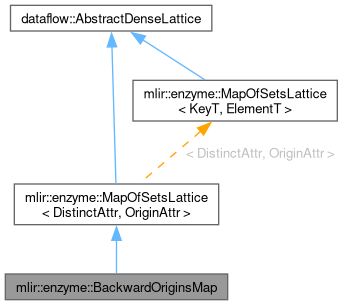
Detailed Description
Definition at line 152 of file ActivityAnnotations.h.
Member Function Documentation
◆ getOrigins()
|
inline |
Definition at line 160 of file ActivityAnnotations.h.
References mlir::enzyme::MapOfSetsLattice< DistinctAttr, OriginAttr >::lookup().
Referenced by mlir::enzyme::DenseBackwardActivityAnnotationAnalysis::visitOperation().
◆ markAllOriginsUnknown()
|
inline |
Definition at line 158 of file ActivityAnnotations.h.
References mlir::enzyme::MapOfSetsLattice< DistinctAttr, OriginAttr >::markAllUnknown().
Referenced by mlir::enzyme::DenseBackwardActivityAnnotationAnalysis::visitCallControlFlowTransfer(), and mlir::enzyme::DenseBackwardActivityAnnotationAnalysis::visitOperation().
◆ meet()
|
inlineoverride |
Definition at line 162 of file ActivityAnnotations.h.
References mlir::enzyme::MapOfSetsLattice< DistinctAttr, OriginAttr >::join().
◆ print()
|
override |
Definition at line 375 of file ActivityAnnotations.cpp.
References printMapOfSetsLattice().
The documentation for this class was generated from the following files:
- MLIR/Analysis/ActivityAnnotations.h
- MLIR/Analysis/ActivityAnnotations.cpp
Generated on Fri May 8 2026 19:56:27 for Enzyme by